Adipose tissue macrophages (ATMs) sit at the intersection of immune surveillance and metabolic regulation. Their crosstalk with insulin signaling is a multi-layered process in which immune activation directly disrupts the molecular machinery of insulin action.
This review systematically delineates the application of single-cell RNA sequencing in characterizing adipose tissue macrophages during insulin resistance, describes the in vitro co-culture systems utilized to investigate macrophage–adipocyte interactions, and examines the production of interleukin‑6 (IL‑6) by adipose tissue macrophages alongside its regulatory impact on insulin receptor signaling. Furthermore, it addresses the crosstalk between macrophage‑derived IL‑1β and the impairment of insulin signaling in adipocytes, and provides an analysis of M1/M2 macrophage polarization markers, including CD86 and CD206, within insulin‑resistant adipose tissue.
Table of Contents
1. Single-cell RNA sequencing of adipose tissue macrophages in insulin resistance
2. Co-culture systems for studying macrophage–adipocyte interactions in vitro
3. IL-6 production from adipose tissue macrophages and its effect on insulin receptor signaling
4. Crosstalk between macrophage-derived IL-1β and insulin signaling impairment in adipocytes
5. Analysis of M1/M2 macrophage polarization markers (CD86, CD206) in insulin-resistant adipose tissue
01 Single-cell RNA sequencing of adipose tissue macrophages in insulin resistance
Single‑cell RNA sequencing (scRNA‑seq) has revolutionized the understanding of adipose tissue macrophage (ATM) heterogeneity, moving beyond the classic M1/M2 polarization model to define functionally distinct subpopulations that directly perturb insulin signaling pathways in obesity and insulin resistance. Specifically, this high‑resolution approach identifies subset‑specific transcriptional signatures, thereby providing mechanistic insights into how distinct ATM populations disrupt key insulin signaling pathways and drive metabolic dysfunction.
A key discovery from scRNA‑seq is lipid‑associated macrophages (LAMs), particularly the Triggering Receptor Expressed on Myeloid Cells 2 (TREM2) subset, which is abundant in obese adipose tissue and dedicated to lipid metabolism[1]. Specifically, scRNA‑seq reveals that LAMs exhibit gene programs governing lipid uptake, processing and storage. For instance, the lysosomal protein TM4SF19 (Transmembrane 4 Superfamily Member 19), selectively expressed in LAMs, suppresses lysosomal acidification, leading to lipid intermediate accumulation, inflammation, and exacerbated insulin resistance through dysregulation of insulin signaling pathways. Furthermore, TREM2+ LAMs are linked to adipose fibrosis and crown‑like structures (CLSs), which directly disrupt adipocyte function and interfere with insulin signaling pathways that are critical for maintaining whole‑body glucose homeostasis[1,2].
scRNA‑seq further identifies CD9+ ATMs as a metabolically activated population distinct from classic CD11c+ (Cluster of Differentiation 11c) M1‑like macrophages. These cells display unique transcriptional profiles associated with inflammatory and metabolic activities, and their surface markers thus represent promising therapeutic targets for restoring insulin signaling pathways. Additionally, TIM4+ (T‑Cell Immunoglobulin and Mucin Domain Containing 4) resident macrophages have been recognized as novel regulators of adiposity, specifically, they modulate adipocyte size via platelet‑derived growth factor‑CC (PDGF‑CC) and remodel adipose tissue architecture, thereby indirectly influencing insulin signaling pathways and insulin sensitivity[2].
Beyond identifying subsets, scRNA‑seq also clarifies how environmental cues such as nutrient overload and hypoxia shape ATM plasticity. Specifically, it has uncovered dysregulated long non‑coding RNAs (lncRNAs) including HEM2ATM (HEM2 Associated Long Non‑Coding RNA in Adipose Tissue Macrophages) in obese ATMs, which promote meta‑inflammation and insulin resistance through interactions with heterogeneous nuclear ribonucleoprotein U (hnRNP U) and subsequent disruption of insulin signaling pathways. Moreover, scRNA‑seq links myeloid G‑protein‑signaling modulator 1 (GPSM1) to metabolic inflammation and type 2 diabetes (T2D), and reveals that targeting IRE1α (Inositol‑Requiring Enzyme 1α) alleviates insulin resistance by suppressing pathogenic ATM subsets and restoring aberrant insulin signaling pathways[3].
Collectively, scRNA‑seq demonstrates that insulin resistance arises from the coordinated actions of specialized ATM subsets rather than generic inflammation. These findings thus support the development of targeted therapies, such as promoting anti‑inflammatory ATM phenotypes using rosiglitazone or large yellow tea polysaccharide (LYTP), and glucocorticoid‑mediated macrophage modulation, all of which aim to restore impaired insulin signaling pathways. Importantly, scRNA‑seq enables precise monitoring of subset‑specific responses, thereby facilitating therapeutic refinement and more effective restoration of insulin signaling pathways in metabolic disease[4].
In conclusion, scRNA‑seq has redefined ATM heterogeneity beyond the M1/M2 framework by identifying LAMs, CD9+, and TIM4+ subpopulations with distinct roles in lipid metabolism, inflammation, fibrosis, and intercellular crosstalk[5]. Ultimately, this high‑resolution insight is critical for designing precision therapies against obesity‑related metabolic diseases, with a central focus on restoring dysregulated insulin signaling pathways.
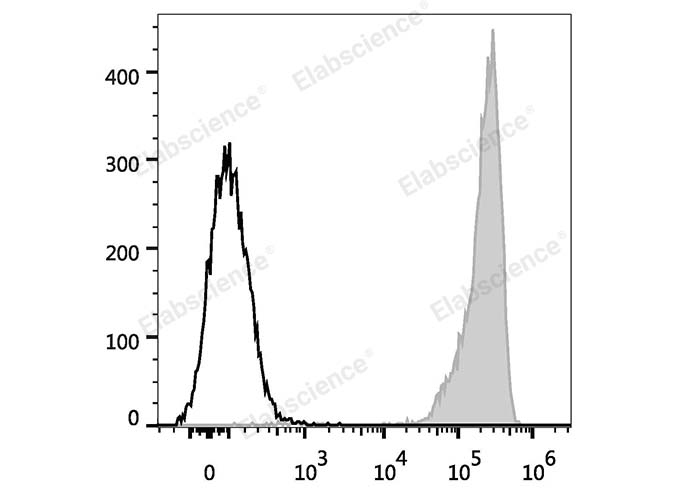
Fig. 1 Analysis of CD9 expression in human platelets. Human platelets are stained with FITC Anti-Human CD9 Antibody (filled gray histogram; cat. no. E-AB-F1086C). Unstained platelets (empty black histogram; cat. no. E-AB-F09792C) served as the control. (The data are provided by Elabscience.)
02 Co-culture systems for studying macrophage–adipocyte interactions in vitro
Co-culture systems are essential for investigating paracrine and inflammatory crosstalk between macrophages and adipocytes in vitro, especially in obesity and type 2 diabetes mellitus (T2DM). These systems recapitulate the microenvironment of adipose tissue, a dynamic endocrine organ whose dysfunction drives systemic insulin resistance and chronic low-grade inflammation. Adipose tissue inflammation, characterized by immune cell infiltration and pro-inflammatory cytokine secretion, is a hallmark of metabolic disease pathogenesis[6].
Diverse co-culture platforms have been established, from conventional two-dimensional (2D) models to more physiologically relevant three-dimensional (3D) systems. These approaches overcome limitations of monocultures and animal models by enabling controlled mechanistic studies[7].
2.1 2D Co-culture Systems
2D co-culture is performed on planar substrates, either via direct cell–cell contact or indirect culture using Transwell porous membranes. In direct co-culture, mixed seeding of adipocytes and macrophages supports both physical contact and paracrine signaling, which is critical for certain inflammatory responses. For example, lipopolysaccharide (LPS)-activated RAW 264.7 macrophages enhance leptin production in 3T3-L1 adipocytes, forming an inflammatory loop. Human macrophages also upregulate adipocyte inflammatory mediators, a response modulated by second-generation antipsychotics and glucocorticoids[8].
In indirect Transwell co-culture, soluble factor exchange occurs without physical contact, making it widely used to study adipokine and cytokine effects. Macrophage-conditioned medium promotes adipocyte-associated breast cancer progression, while diosmetin and passion fruit seed extract attenuate inflammation and lipolysis by suppressing nitric oxide (NO), tumor necrosis factor-α (TNF-α), p38 mitogen-activated protein kinase (p38 MAPK), and nuclear factor-κB (NF-κB) signaling[9].
Although 2D systems are simple and suitable for high-throughput screening (HTS), they poorly replicate the 3D architecture and mechanical cues of native adipose tissue.
2.2 3D Co-culture Systems
3D models were developed to better mimic the in vivo adipose microenvironment and support more physiological adipogenesis and cell interactions.Spheroid models enable self-assembled tissue-like structures and multi-cellular crosstalk. Studies using 3T3-L1 preadipocytes, J774 macrophages, and fibroblasts demonstrate spontaneous adipogenic differentiation in this context.Scaffold-based systems provide tunable structural support, while scaffold-free 3D culture under chemically defined conditions improves adipocyte maturation.Microfluidic and organ-on-a-chip platforms allow precise control over nutrient and oxygen gradients and dynamic cell interactions. For instance, adipocyte-derived lipids promote cancer cell migration via hypoxia-inducible factor-1α (HIF-1α) activation in metastasis-on-a-chip models, with porous membranes facilitating physiological crosstalk[10,11].
2.3 Animal-Free and Humanized Models
Ethical concerns and poor interspecies translatability have driven the development of animal-free co-culture systems, which replace fetal bovine serum (FBS) with human alternatives such as human platelet lysate (hPL). A 3D animal-free adipocyte–macrophage co-culture has been established to study inflammation in vitro, offering a more clinically relevant platform for obesity-related drug testing[12].
Advanced co-culture systems, including organoids and microfluidic models, continue to improve physiological relevance. Incorporation of endothelial cells, fibroblasts, and other stromal components further recapitulates adipose tissue complexity. The shift toward animal-free, human-derived models is critical for translating mechanistic insights into therapies targeting adipocyte–macrophage crosstalk to treat obesity-associated metabolic complications[13].
Fig. 2 Characterization of M1-polarized pro-inflammatory RAW 264.7 macrophages. RAW 264.7 cells were subjected to M1 polarization using RAW 264.7 Polarized M1 Macrophage Induction and Identification Kit (Cat. no. XJM004). Compared with untreated control cells, M1-polarized macrophages exhibited short and dense pseudopodia, markedly increased intracellular granules, and elevated surface expression of F4/80, CD11b and CD86 as determined by flow cytometry. (The data are provided by Elabscience.)
03 IL-6 production from adipose tissue macrophages and its effect on insulin receptor signaling
Interleukin-6 (IL-6) derived from adipose tissue macrophages (ATMs) plays a critical and context-dependent role in regulating insulin signaling, particularly in obesity-associated insulin resistance. As a dynamic endocrine organ, adipose tissue undergoes chronic low-grade meta-inflammation in obesity. This process is characterized by enhanced macrophage infiltration and elevated pro-inflammatory cytokine secretion, including macrophage derived cytokines, and M1-polarized ATMs constitute a major source of IL-6 under obese conditions. The concentration of IL-6 in adipose tissue and systemic circulation can be quantitatively measured using IL-6 ELISA, a reliable method for assessing IL-6 expression levels in metabolic research[14].
Hypertrophic and stressed adipocytes release inflammatory mediators and free fatty acids. These factors promote macrophage polarization toward a pro-inflammatory M1 phenotype. Activated M1 macrophages subsequently secrete TNF-α, IL-1β, and IL-6 as key macrophage derived cytokines, which drive local and systemic inflammation and directly impair insulin signaling[15].
IL-6 acts through two distinct signaling pathways. Classical signaling occurs via membrane-bound IL-6R and gp130, while trans-signaling takes place through soluble IL-6R (sIL-6R) that activates gp130 on cells lacking membrane‑bound IL‑6R. Both of these pathways modulate insulin sensitivity[16].
Mechanistically, elevated IL-6 promotes insulin resistance by disrupting the canonical insulin signaling cascade. Under physiological conditions, insulin induces tyrosine phosphorylation of IRS-1. This event is followed by PI3K/Akt activation and GLUT4 translocation to facilitate glucose uptake[17]. In contrast, IL-6 and TNF-α activate JNK and IKK. These kinases trigger inhibitory serine phosphorylation of IRS‑1 and block downstream signaling. In addition, IL‑6 upregulates SOCS3, which binds the insulin receptor or IRS‑1 to promote its ubiquitination and degradation, further inhibiting PI3K/Akt signaling. Moreover, IL‑6 enhances hepatic gluconeogenesis and blunts insulin‑mediated suppression of glycogenolysis, thereby exacerbating systemic glucose dysregulation. Changes in IL-6 expression during these processes can be accurately quantified by IL-6 ELISA, providing direct evidence for its role in insulin resistance[18].
IL‑6 trans‑signaling also contributes to macrophage recruitment into adipose tissue. Although trans‑signaling blockade reduces ATM infiltration in mice, this intervention does not consistently improve insulin resistance. This observation highlights the context‑dependent and paradoxical actions of IL-6. For example, exercise‑induced IL‑6 enhances insulin sensitivity, which is in contrast to its pro‑inflammatory effects in obesity, a difference that can be verified by comparing IL-6 levels via IL-6 ELISA in different physiological states[19].
Adipocytes also secrete IL‑6. This adipocyte‑derived IL‑6 sensitizes macrophages to IL‑4 signaling and mediates bidirectional crosstalk between adipocytes and immune cells. Furthermore, IL‑1β and TNF-α synergistically increase adipocyte IL‑6 expression, further enriching the pool of macrophage derived cytokines and adipocyte-derived IL-6 in adipose tissue. This interaction establishes a self‑sustaining inflammatory cycle that amplifies insulin resistance[20].
Advanced human in vitro vascularized adipose tissue models are critical for clarifying these complex interactions and translating preclinical findings to human physiology. Since meta‑inflammation is a key driver of insulin resistance and type 2 diabetes, targeting IL‑6 signaling, especially trans‑signaling, represents a promising therapeutic strategy. However, the overall metabolic impact of such interventions requires careful and context‑dependent evaluation, with IL-6 ELISA Kit serving as a valuable tool to monitor changes in IL-6 levels and assess therapeutic efficacy[21].
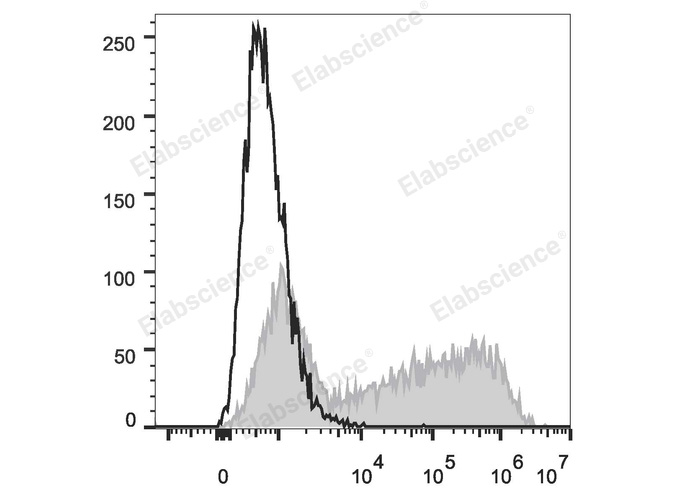
Fig. 3 Analysis of IL-6 expression in HEK293T cells. HEK293T cells were transiently transfected with a pcDNA3.1 plasmid encoding the human IL-6 gene, then stained with APC Anti-Human IL-6 Antibody (filled gray histogram, cat. no. E-AB-F1206E) or APC Rat IgG1, κ Isotype Control (empty black histogram, cat. no. E-AB-F09822E). (The data are provided by Elabscience.)
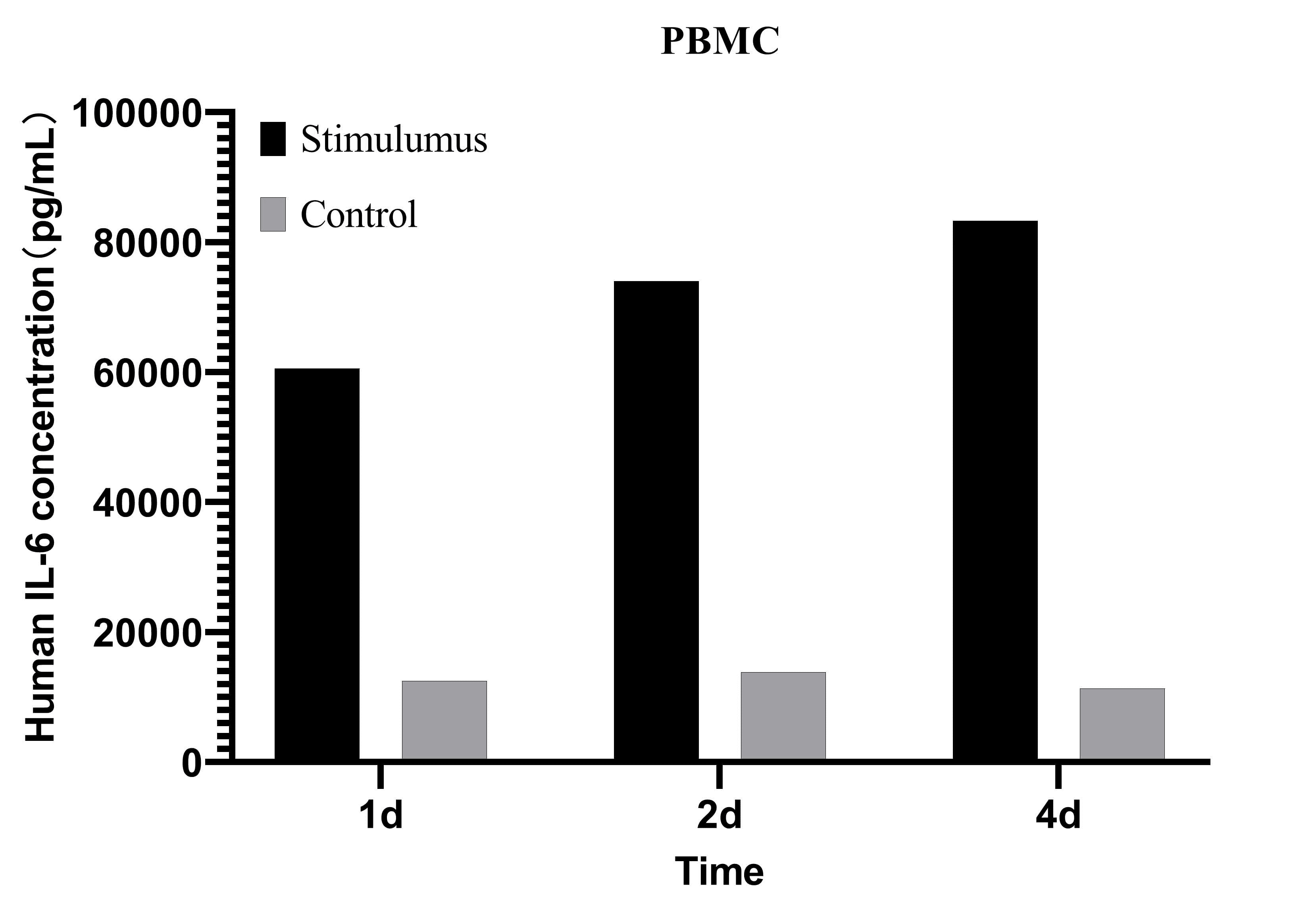
Fig. 4 Analysis of IL-6 levels in the culture supernatant of human PBMCs. Human PBMCs were stimulated with 2 μg/mL PHA for 1, 2, and 4 days. IL-6 concentrations in the culture supernatant were measured using the CellaQuant™ Human IL-6 (Interleukin 6) ELISA Kit (Cat. no. CQH001), with unstimulated cells as the control. (The data are provided by Elabscience.)
04 Crosstalk between macrophage-derived IL-1β and insulin signaling impairment in adipocytes
The intricate crosstalk between macrophage-derived interleukin-1 beta (IL-1β) and adipocyte insulin signaling impairment is a critical mechanism in the pathogenesis of obesity-induced insulin resistance and type 2 diabetes (T2D). Adipose tissue, not merely a passive energy storage depot, functions as an active endocrine organ housing diverse cell types, including adipocytes, pre-adipocytes, and immune cells, predominantly macrophages. In obesity, adipocyte hypertrophy and subsequent hypoxic conditions within adipose tissue trigger a pro-inflammatory cascade that promotes macrophage recruitment and activation, driving their polarization toward a pro-inflammatory M1 phenotype. These activated M1 macrophages are key producers of inflammatory cytokines, including macrophage derived cytokines, with IL-1β playing a prominent role in mediating detrimental effects on adipocyte function and insulin sensitivity[22,23].
As a potent pro-inflammatory cytokine, IL-1β is involved in numerous inflammatory conditions and plays a major role in immune system signal transduction. Within adipose tissue, its expression is significantly upregulated in obesity, and the level of IL-1β can be quantitatively detected using IL-1β ELISA Kit. Macrophage-derived factors, particularly macrophage derived cytokines, markedly alter adipocyte function by inducing inflammatory responses and reducing insulin sensitivity, with IL-1β specifically mediating macrophage-induced impairment of insulin signaling in human primary adipocytes. This cytokine is primarily produced in monocytes and macrophages, with minimal production in adipocytes. Its active form is generated by caspase-1-mediated cleavage of the inactive precursor pro-IL-1β, and caspase-1 is activated by the NOD-like receptor family pyrin domain-containing 3 (NLRP3) inflammasome complex. Transcriptome analysis of human adipocytes further highlights the involvement of the NOD-like receptor pathway, including NLRP3 and PYCARD, in obesity-induced adipose inflammation and the regulation of caspase-1-mediated IL-1β processing[24].
The mechanism by which IL-1β impairs adipocyte insulin signaling is multifaceted. Upon binding to its receptor (IL-1R1) on adipocytes, IL-1β triggers downstream activation of key inflammatory pathways, including nuclear factor-kappa B (NF-κB) and c-Jun N-terminal kinase (JNK). As a master regulator of inflammatory responses, NF-κB activation induces inhibitory serine phosphorylation of insulin receptor substrate-1 (IRS-1), particularly at Ser307. This serine phosphorylation disrupts IRS-1 tyrosine phosphorylation, an essential step for initiating the downstream phosphatidylinositol 3-kinase (PI3K)-Akt pathway and the core axis of insulin action, thereby compromising the entire insulin signaling cascade and reducing adipocyte glucose uptake and utilization[25].
Beyond direct interference with IRS-1, IL-1β contributes to insulin resistance through additional mechanisms. It suppresses the translocation of glucose transporter type 4 (GLUT4) to the cell surface, further limiting adipocyte glucose uptake. IL-1β also promotes lipolysis (fat breakdown), which increases circulating free fatty acids, induces lipotoxicity in various tissues, and exacerbates insulin resistance. Additionally, it reduces the secretion of adiponectin, an anti-inflammatory adipokine that normally enhances insulin sensitivity, further shifting the balance toward an inflammatory and insulin-resistant state. This IL-1β-mediated crosstalk between macrophages and adipocytes, along with other macrophage derived cytokines, is a key component of obesity- and T2D-associated meta-inflammation[26].
Clinical evidence supports the role of IL-1β in insulin resistance. Blocking IL-1 with the interleukin-1 receptor antagonist anakinra improves first-phase insulin secretion and the insulinogenic index in individuals with impaired glucose tolerance, suggesting that targeting IL-1β pathways could be a potential therapeutic strategy for enhancing insulin sensitivity and managing T2D. For instance, 14-Deoxygarcinol improves insulin sensitivity by mitigating NF-κB/Sirtuin 2-NLRP3-mediated adipose tissue remodeling and reducing IL-1β production, which can be validated by IL-1β ELISA Kit. Similarly, puerarin attenuates insulin resistance by suppressing inflammation through modulation of the JNK and IKKβ/NF-κB pathways in adipose tissue. Furthermore, high endogenously synthesized n-3 polyunsaturated fatty acids attenuate high-fat diet-induced insulin resistance by inhibiting NLRP3 inflammasome activation via the Akt/GSK-3β/TXNIP pathway, thereby regulating IL-1β secretion and the release of other macrophage derived cytokines[27,28].
Adipose tissue inflammation and its impact on insulin signaling are well documented. Obesity promotes adipose tissue expansion, leading to immune cell infiltration, most notably by macrophages, which adopt a pro-inflammatory phenotype and secrete key cytokines such as TNF-α and IL-1β. Accordingly, continuous development of compounds that specifically target the IL-1β pathway, its upstream activators, or downstream effectors holds promise for establishing effective treatments for insulin resistance and its associated metabolic disorders.
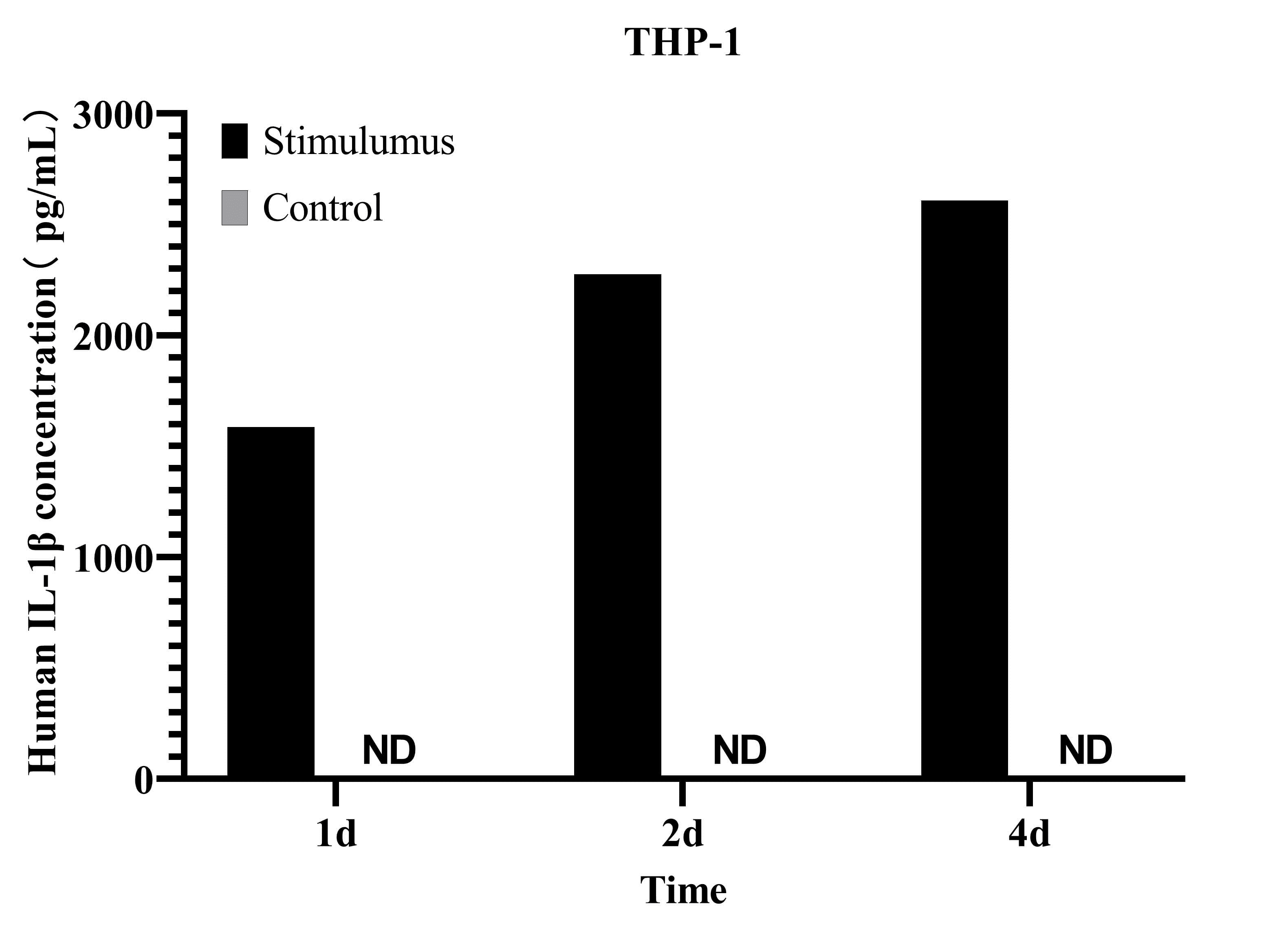
Fig. 5 Analysis of IL-1β levels in the culture supernatant of THP-1 cells. THP-1 cells were stimulated with 100 ng/mL PMA and 500 ng/mL LPS for 1, 2, and 4 days. IL-1β concentrations in the culture supernatant were measured using the CellaQuant™ Human IL-1β (Interleukin 1 Beta) ELISA Kit (Cat. no. CQH015), with unstimulated cells as the control. (The data are provided by Elabscience.)
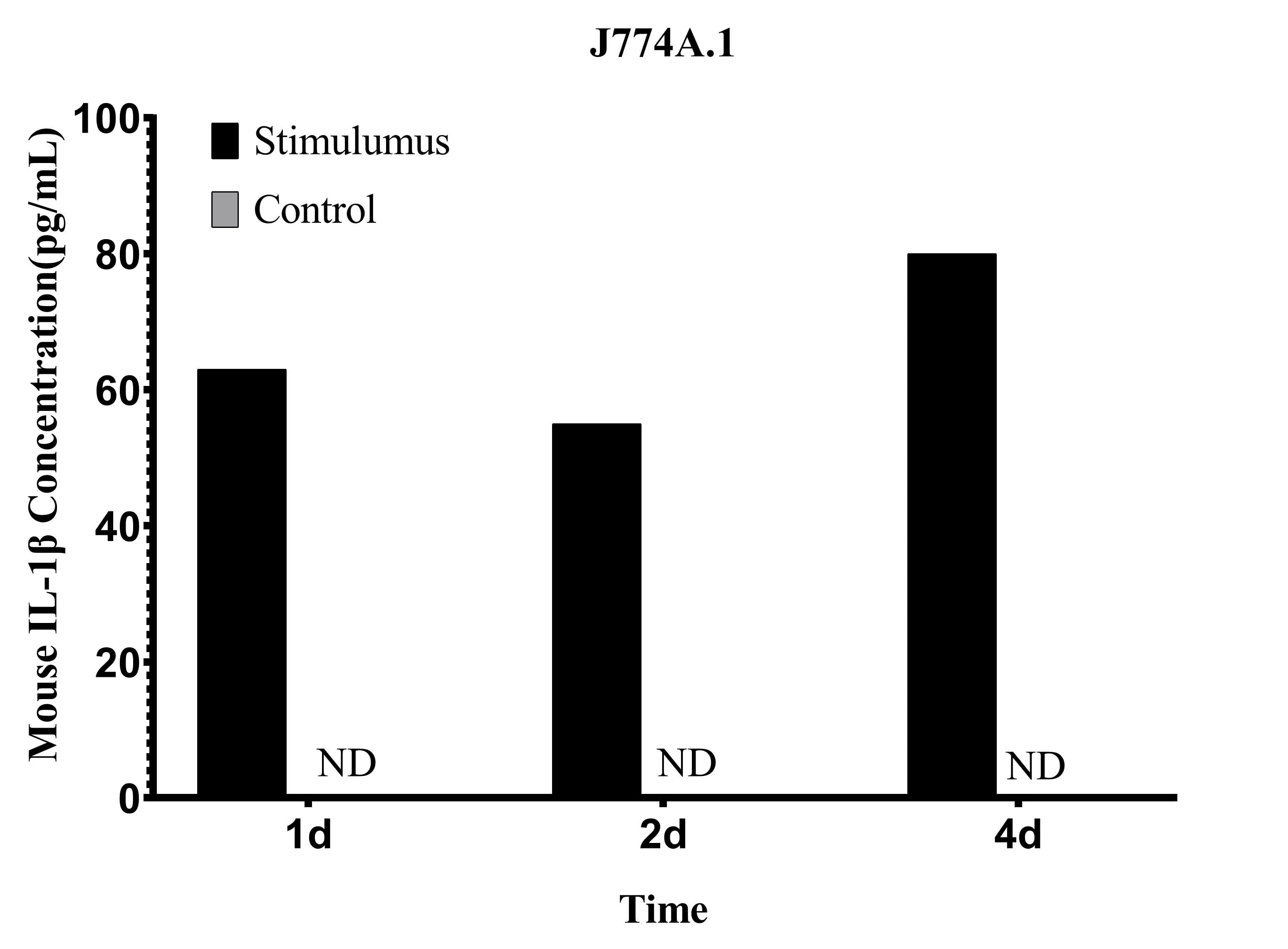
Fig. 6 Analysis of IL-1β levels in the culture supernatant of J774A.1 cells. J774A.1 cells were stimulated with 1 μg/mL LPS for 1, 2, and 4 days. IL-1β concentrations in the culture supernatant were measured using the CellaQuant™ Mouse IL-1β (Interleukin 1 Beta) ELISA Kit (Cat. no. CQM003), with unstimulated cells as the control. (The data are provided by Elabscience.)
05 Analysis of M1/M2 macrophage polarization markers (CD86, CD206) in insulin-resistant adipose tissue
Macrophage polarization within adipose tissue, particularly the M1 and M2 phenotypes, is a critical determinant of local inflammation and systemic insulin resistance in obesity. M1 macrophages, characterized by markers such as CD86, are pro-inflammatory and contribute to insulin resistance, whereas M2 macrophages, marked by CD206, are anti-inflammatory and generally associated with insulin sensitivity and tissue repair. The balance between these two phenotypes is significantly altered in insulin-resistant adipose tissue[29].
In obese individuals and animal models of diet-induced obesity, there is a distinct shift in adipose tissue macrophage (ATM) populations toward a pro-inflammatory M1 phenotype. This M1 polarization is driven by signals from hypertrophied and stressed adipocytes, which release inflammatory mediators including saturated free fatty acids. These signals activate Toll-like Receptor 4 (TLR4) on macrophages, leading to the activation of NF-κB and subsequent production of pro-inflammatory M1 mediators, such as tumor necrosis factor-alpha (TNF-α), interleukin-1β (IL-1β), and nitric oxide (NO) via inducible Nitric Oxide Synthase (iNOS). These M1 macrophages interfere with insulin signaling in adjacent cells, thereby directly contributing to the development of systemic insulin resistance and type 2 diabetes[30,31].
CD86, a costimulatory molecule, is a well-recognized marker for M1 polarized macrophages, and its detection can be achieved using an anti CD86 antibody. CD86 expression in adipose tissue is increased in obesity and insulin resistance. For instance, in rats, diabetes induces modulation of hepatic CD86 expression, which is associated with insulin resistance and inflammation. The upregulation of CD86+ M1 polarized macrophages correlates with elevated TNF-α and IL-6 expression, indicative of a pro-inflammatory state. Interferon Regulatory Factor (IRF)-5, a molecule involved in M1 macrophage polarization, also exhibits increased adipose tissue expression in obesity, further linking M1 activation to metabolic inflammation[32,33].
Conversely, M2 polarized macrophages are characterized by the expression of CD206, also known as the mannose receptor, and an anti CD206 antibody is commonly used to identify this macrophage subtype. These M2 polarized macrophages possess anti-inflammatory properties, secrete cytokines such as IL-10 and Transforming Growth Factor-beta (TGF-β), and participate in tissue repair, wound healing, and inflammation resolution. In lean individuals, M2 ATMs constitute the majority of adipose tissue macrophages. In obesity and insulin resistance, however, the proportion of CD206+ M2 polarized macrophages tends to decrease or their function becomes impaired. While the classical paradigm suggests reduced M2 polarized macrophages, some human obese adipose tissue studies have reported that despite increased total macrophage numbers, these cells might exhibit an anti-inflammatory profile even with enhanced CD206 expression. For example, a pro-inflammatory CD11c+CD206+ ATM subset has been identified in human obesity and is closely associated with insulin resistance, suggesting a more complex, heterogeneous macrophage response that extends beyond a simple M1/M2 dichotomy[34].
The spatial distribution of macrophages also plays a crucial role in adipose tissue inflammation and insulin resistance. In insulin-resistant adipose tissue, M1 polarized macrophages often form crown-like structures (CLS) around dead or dying adipocytes. These CLS are considered a hallmark of adipose tissue inflammation and are predominantly composed of CD11c-positive M1 polarized macrophages. This localized inflammation within adipose tissue significantly contributes to systemic insulin resistance[35].
Furthermore, the microenvironment within adipose tissue can modulate macrophage polarization, shifting the balance between M1 polarized macrophages and M2 polarized macrophages. Adipocyte-derived microvesicles from obese mice have been shown to induce an M1 macrophage phenotype through secreted miR-155, highlighting a key mechanism by which adipocytes promote inflammation. Similarly, adipocyte-derived exosomes carrying Sonic Hedgehog mediate M1 macrophage polarization-induced insulin resistance via Ptch and PI3K pathways. Conversely, telmisartan, an angiotensin II type 1 receptor blocker, improves insulin resistance and modulates ATM polarization toward a more anti-inflammatory state, favoring M2 polarized macrophages in high-fat-fed mice. Exercise interventions also induce a phenotypic switch in ATM polarization, reducing the M1 pro-inflammatory state and promoting M2 polarized macrophages to improve insulin sensitivity[36].
The interaction between different macrophage subtypes is another critical factor. Partial depletion of CD206+ M2-like macrophages affects the proliferation of beige progenitors and enhances browning after cold stimulation, indicating a role for M2 macrophages in thermogenesis and energy expenditure. Additionally, CD226+ adipose tissue macrophages, derived from monocyte precursors, have been implicated in the regulation of lipid metabolism[37,38].
In summary, the precise characterization of macrophage polarization in insulin-resistant adipose tissue involves assessing markers such as CD86 for M1 polarized macrophages (detectable via anti CD86 antibody) and CD206 for M2 polarized macrophages (identifiable using anti CD206 antibody). The shift toward increased CD86 expression, alongside potentially altered CD206 expression and the formation of crown-like structures, collectively indicates a pro-inflammatory state that underpins the development of insulin resistance in obesity. Therefore, understanding these dynamics is crucial for developing therapeutic strategies targeting adipose tissue inflammation and insulin resistance.
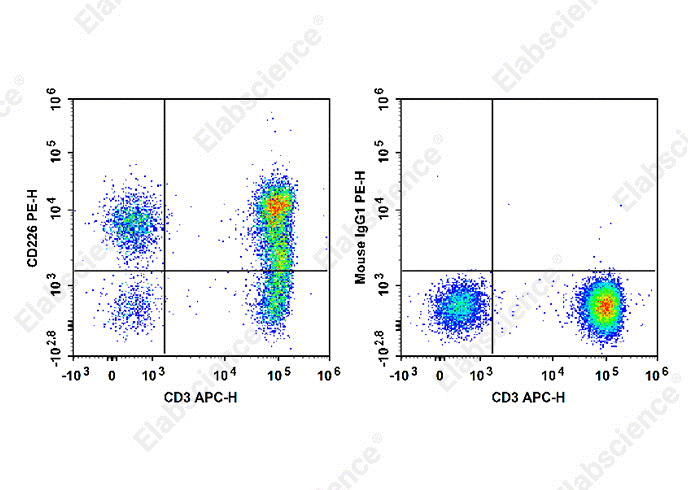
Fig. 7 Analysis of CD226 expression in human peripheral blood cells. Normal human peripheral blood cells were stained with APC Anti-Human CD3 Antibody [OKT-3] (Cat. no. E-AB-F1001E) together with PE Anti-Human CD226/DNAM-1 Antibody [11A8] (Cat. no. E-AB-F1369D) (left panel) or PE Mouse IgG1, κ Isotype Control (Cat. no. E-AB-F09792D) (right panel). Analysis was performed on cells within the lymphocyte gate. (The data are provided by Elabscience.)
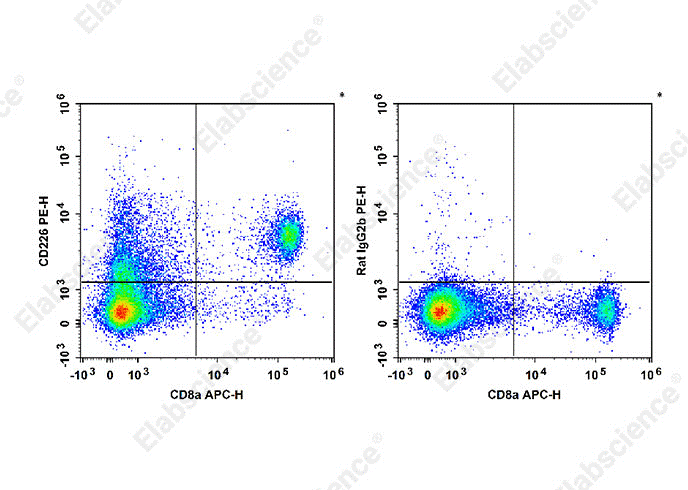
Fig. 8 Analysis of CD226 expression in C57BL/6 mouse splenocytes. Normal C57BL/6 splenocytes were stained with APC Anti-Mouse CD8a Antibody (Cat. no. E-AB-F1104E) together with PE Anti-Mouse CD226 Antibody [10E5] (Cat. no. E-AB-F1402D) (left panel) or PE Rat IgG2b, κ Isotype Control (Cat. no. E-AB-F09842D) (right panel). Analysis was performed on cells within the total viable cell gate. (The data are provided by Elabscience.)
References:
[1] Yan X, Miok V, Karlina R, et al. 1666-P: ScRNAseq–Based Analysis of the Stromal Immune Cell Composition in Murine Preweaning and Adult White Adipose Tissues[J]. Diabetes, 2023, 72(Supplement_1): 1666-P.
[2] Huh J Y, Kim J B. TIM4+ adipose tissue-resident macrophages: new modulators of adiposity[J]. Nature Reviews Endocrinology, 2021, 17(11): 645-646.
[3] Vijay J, Gauthier M F, Biswell R L, et al. Single-cell analysis of human adipose tissue identifies depot-and disease-specific cell types[J]. Nature metabolism, 2020, 2(1): 97-109.
[4] Wu H, Kong X, Shan R, et al. Eurotium cristatum-Fermented White Tea Ameliorates DSS-Induced Colitis by Multi-Scale[J]. Foods, 2025, 15(1): 72.
[5] Magalhaes M S, Smith P, Portman J R, et al. Role of Tim4 in the regulation of ABCA1+ adipose tissue macrophages and post-prandial cholesterol levels[J]. Nature communications, 2021, 12(1): 4434.
[6] Monk J M, Liddle D M, Hutchinson A L, et al. Studying adipocyte and immune cell cross talk using a co-culture system[M]//Immunometabolism: Methods and Protocols. New York, NY: Springer US, 2020: 111-130.
[7] Seo J, Kim K S, Park J W, et al. Metastasis-on-a-chip reveals adipocyte-derived lipids trigger cancer cell migration via HIF-1α activation in cancer cells[J]. Biomaterials, 2021, 269: 120622.
[8] Caldari-Torres C, Beck J. Effects of co-incubation of LPS-stimulated RAW 264.7 macrophages on leptin production by 3T3-L1 adipocytes: a method for co-incubating distinct adipose tissue cell lines[J]. Bulletin of the National Research Centre, 2022, 46(1): 57.
[9] Lee H, Sung J, Kim Y, et al. Inhibitory effect of diosmetin on inflammation and lipolysis in coculture of adipocytes and macrophages[J]. Journal of food biochemistry, 2020, 44(7): e13261.
[10] Nowakowski S, Nellinger S, Albrecht F B, et al. Animal‐free setup of a 3D mature adipocyte macrophage co‐culture to induce inflammation in vitro[J]. Advanced healthcare materials, 2025, 14(22): 2500779.
[11] Shen J X, Couchet M, Dufau J, et al. 3D adipose tissue culture links the organotypic microenvironment to improved adipogenesis[J]. Advanced Science, 2021, 8(16): 2100106.
[12] Mora I, Puiggros F, Serras F, et al. Emerging models for studying adipose tissue metabolism[J]. Biochemical Pharmacology, 2024, 223: 116123.
[13] Orabi M, Yeganeh M S, Chun T H, et al. Modeling Adipokine and Insulin‐Mediated Crosstalk Between Adipocytes and Beta Cells Using Flow‐Enabled Microfluidics[J]. Small, 2025, 21(35): 2504686.
[14] Boutens L, Hooiveld G J, Dhingra S, et al. Unique metabolic activation of adipose tissue macrophages in obesity promotes inflammatory responses[J]. Diabetologia, 2018, 61(4): 942-953.
[15] Tessaro F H G, Ayala T S, Nolasco E L, et al. Insulin influences LPS-induced TNF-α and IL-6 release through distinct pathways in mouse macrophages from different compartments[J]. Cellular Physiology and Biochemistry, 2017, 42(5): 2093-2104.
[16] Carey A L, Febbraio M A. Interleukin-6 and insulin sensitivity: friend or foe?[J]. Diabetologia, 2004, 47(7): 1135-1142.
[17] Sayem A S M, Arya A, Karimian H, et al. Action of phytochemicals on insulin signaling pathways accelerating glucose transporter (GLUT4) protein translocation[J]. Molecules, 2018, 23(2): 258.
[18] Park J E, Han J S. Improving the effect of ferulic acid on inflammation and insulin resistance by regulating the JNK/ERK and NF-κB pathways in TNF-α-treated 3T3-L1 adipocytes[J]. Nutrients, 2024, 16(2): 294.
[19] Kraakman M J, Kammoun H L, Allen T L, et al. Blocking IL-6 trans-signaling prevents high-fat diet-induced adipose tissue macrophage recruitment but does not improve insulin resistance[J]. Cell metabolism, 2015, 21(3): 403-416.
[20] Aguree S. Iron–Inflammasome Crosstalk in Adipose Tissue: Unresolved Roles of NLRP3 and IL-1β in Metabolic Inflammation[J]. International Journal of Molecular Sciences, 2025, 26(17): 8304.
[21] Akbari M, Hassan-Zadeh V. IL-6 signalling pathways and the development of type 2 diabetes[J]. Inflammopharmacology, 2018, 26(3): 685-698.
[22] Garg S S, Kushwaha K, Dubey R, et al. Association between obesity, inflammation and insulin resistance: Insights into signaling pathways and therapeutic interventions[J]. Diabetes research and clinical practice, 2023, 200: 110691.
[23] Ghemiș L, Goriuc A, Minea B, et al. Myeloid-derived suppressor cells (MDSCs) and obesity-induced inflammation in type 2 diabetes[J]. Diagnostics, 2024, 14(21): 2453.
[24] Chen J, Feng Z, Zhou F, et al. 14-Deoxygarcinol improves insulin sensitivity in high-fat diet-induced obese mice via mitigating NF-κB/Sirtuin 2-NLRP3-mediated adipose tissue remodeling[J]. Acta Pharmacologica Sinica, 2023, 44(2): 434-445.
[25] Wang K, Li F, Cui Y, et al. The association between depression and type 1 diabetes mellitus: inflammatory cytokines as ferrymen in between?[J]. Mediators of inflammation, 2019, 2019(1): 2987901.
[26] Acosta-Martinez M, Cabail M Z. The PI3K/Akt pathway in meta-inflammation[J]. International journal of molecular sciences, 2022, 23(23): 15330.
[27] Gandhi G R, Hillary V E, Antony P J, et al. A systematic review on anti-diabetic plant essential oil compounds: Dietary sources, effects, molecular mechanisms, and safety[J]. Critical reviews in food science and nutrition, 2024, 64(19): 6526-6545.
[28] Yan S, Santoro A, Niphakis M J, et al. Inflammation causes insulin resistance in mice via interferon regulatory factor 3 (IRF3)-mediated reduction in FAHFA levels[J]. Nature Communications, 2024, 15(1): 4605.
[29] Boutens L, Stienstra R. Adipose tissue macrophages: going off track during obesity[J]. Diabetologia, 2016, 59(5): 879-894.
[30] Chylikova J, Dvorackova J, Tauber Z, et al. M1/M2 macrophage polarization in human obese adipose tissue[J]. Biomed Pap Med Fac Univ Palacky Olomouc Czech Repub, 2018, 162(2): 79-82.
[31] Saitoh S, Van Wijk K, Nakajima O. Crosstalk between metabolic disorders and immune cells[J]. International Journal of Molecular Sciences, 2021, 22(18): 10017.
[32] Sheikh R, Shakerian S, Tabatabaei S R F, et al. Moderate and high-intensity interval training protect against diabetes-induced modulation of hepatic CD86 and CD206 expression associated with the amelioration of insulin resistance and inflammation in rats[J]. Immunobiology, 2023, 228(6): 152745.
[33] Sindhu S, Thomas R, Kochumon S, et al. Increased adipose tissue expression of interferon regulatory factor (IRF)-5 in obesity: association with metabolic inflammation[J]. Cells, 2019, 8(11): 1418.
[34] Wentworth J M, Naselli G, Brown W A, et al. Pro-inflammatory CD11c+ CD206+ adipose tissue macrophages are associated with insulin resistance in human obesity[J]. Diabetes, 2010, 59(7): 1648-1656.
[35] Faria S S, Corrêa L H, Heyn G S, et al. Obesity and breast cancer: the role of crown-like structures in breast adipose tissue in tumor progression, prognosis, and therapy[J]. Journal of breast cancer, 2020, 23(3): 233.
[36] Zhang Y, Mei H, Chang X, et al. Adipocyte-derived microvesicles from obese mice induce M1 macrophage phenotype through secreted miR-155[J]. Journal of molecular cell biology, 2016, 8(6): 505-517.
[37] Igarashi Y, Nawaz A, Kado T, et al. Partial depletion of CD206-positive M2-like macrophages induces proliferation of beige progenitors and enhances browning after cold stimulation[J]. Scientific Reports, 2018, 8(1): 14567.
[38] Gallerand A, Caillot Z, Terekhova M, et al. CD226+ adipose tissue macrophages arise from MDP-derived monocytes and regulate lipid metabolism[J]. bioRxiv, 2024: 2024.12. 03.626330.



